Publications
Group highlights
At the end of this page, you can find the full list of publications and patents. All papers are also available on arXiv.
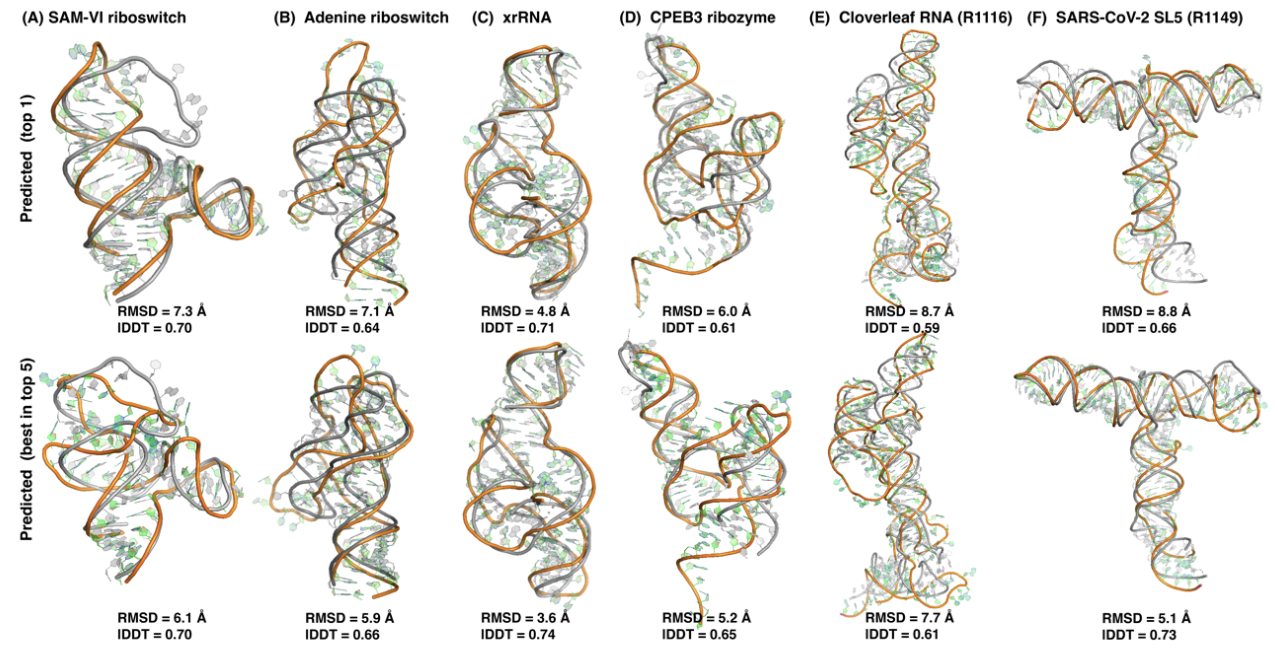
Here, we coupled deep mutational scanning with mobility-based selection to separate structurally stable from nonstable RNA variants. The subsequent high-throughput sequencing allows the detection of the covariational signals of key secondary and tertiary base pairs with significantly improved, RNA-specific, unsupervised analysis of covariation-induced deviation of activity (CODA2) and amplification of these signals by Monte-Carlo simulated annealing. The resulting base-pairing structures can serve as the restraints for final energy-based structure predictions. This MobiSeq technique was tested on four structurally different RNAs. The results show that the detection of tertiary base pairs by MobiSeq allows reasonably accurate prediction of RNA tertiary structures. The proposed method should be useful for structural inference and improved mechanistic understanding of all RNAs with detectable changes in mobility due to deleterious mutations.
Jinle Tang+, Yazhou Shi+, Zhe Zhang, Dailin Luo, Jian Zhan, and Yaoqi Zhou
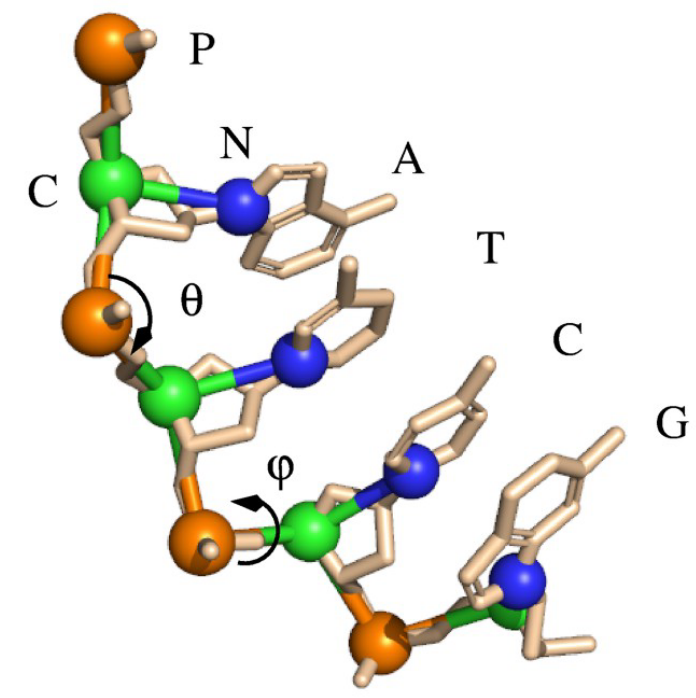
In this work, we present acoarse-grained (CG) model for DNA based on the RNA CG model proposed by us, topredict 3D structures and stability for both dsDNA and ssDNA from the sequence. Combined with aMonte Carlo simulated annealing algorithm and CG force fields involving the sequence-dependentbase-pairing /stacking interactions and an implicit electrostatic potential, the present model successfully folds 20 dsDNAs (�52nt) and 20 ssDNAs (�74nt) into the corresponding native-like structures just from their sequences, with an overall mean RMSD of3.4Å from the experimental structures. For DNAs with various lengths and sequences, the present model can make reliable predictions on stability.
Zi-Chun Mu, Ya-Lan Tan, Ben-Gong Zhang,J ie Liu, Ya-Zhou Shi
PLoS Comput Biol18(10), e1010501 (2022).
See News by YZ Shi.
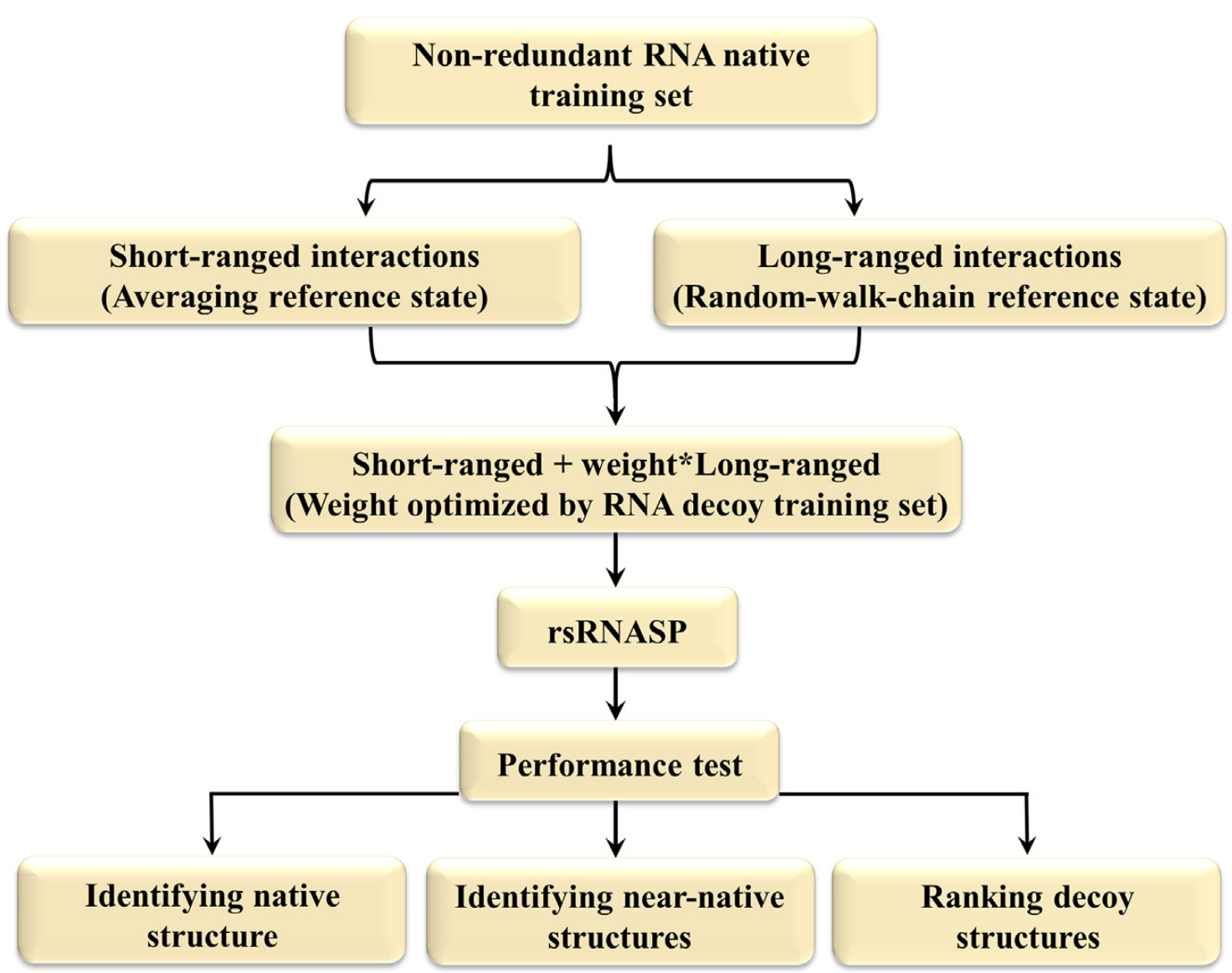
In this work, we have developed an all-atom distance-dependent statistical potential based on residue separation for RNA 3D structure evaluation, namely rsRNASP, which is composed of short- and long-ranged potentials distinguished by residue separation. The extensive examinations against available RNA test datasets show that rsRNASP has apparently higher performance than the existing statistical potentials for the realistic test datasets with large RNAs from structure prediction models, including the newly released RNA-Puzzles dataset, and is comparable to the existing top statistical potentials for the test datasets with small RNAs or near-native decoys..
Ya-Lan Tan, Xunxun Wang, Ya-Zhou Shi, Wenbing Zhang, and Zhi-Jie Tan
Biophysical Journal 21, 142–156 (2022)
See reports
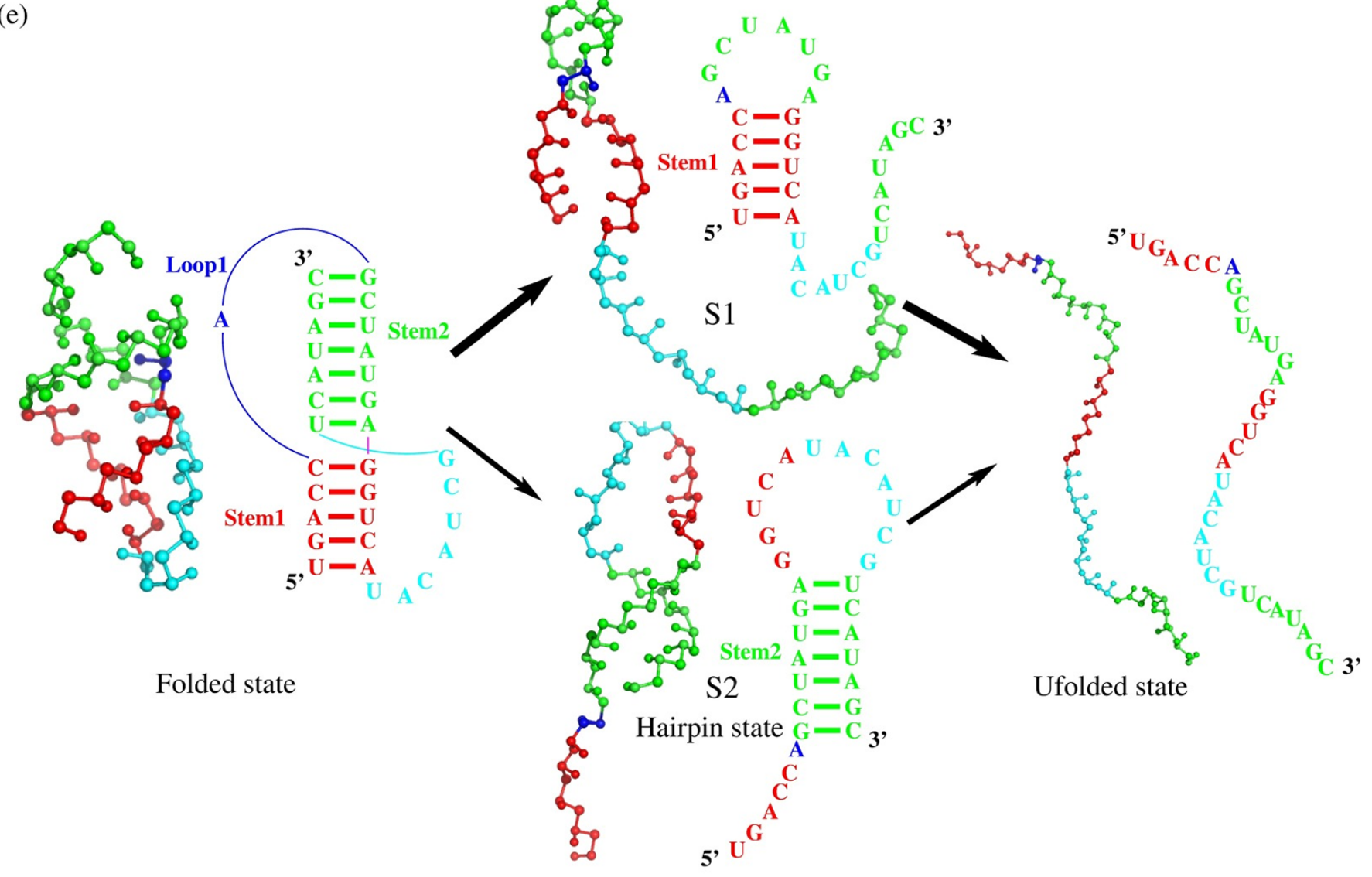
In the work, we employed our previously developed coarse-grained model with implicit salt to make extensive predictions and comprehensive analyses on the 3D structures and stability for RNA pseudoknots in monovalent/divalent ion solutions. The comparisons with available experimental data show that our model can successfully predict the 3D structures of RNA pseudoknots from their sequences, and can also make reliable predictions for the stability of RNA pseudoknots with different lengths and sequences over a wide range of monovalent/divalent ion concentrations. Furthermore, we made comprehensive analyses on the unfolding pathway for various RNA pseudoknots in ion solutions.
Ya-Zhou Shi,Lei Jin,Chen-Jie Feng,Ya-Lan Tan,Zhi-Jie Tan
PLoS Comput Biol14(6) e1006222.(2019)
Patents
Ya-Zhou Shi
RNA 3D structure prediction
PCT/NL20-20/050797 (2025)
Full List of publications
RNA-MobiSeq: Deep mutational scanning and mobility-based selection for RNA base-pairing and tertiary structure inference
Jinle Tang+, Yazhou Shi+, Zhe Zhang, Dailin Luo, Jian Zhan, and Yaoqi Zhou
BioRxiv (2024)
TiRNA: a coarse-grained method with temperature and ion effects for RNA structure folding and prediction
Wang X, Zhou Z, Tan, Y.-L, Shi, Y.-Z, and Tan, Z.-J*.
Nucleic Acids Research (2026)
R3J-AGNN: GNN-Based Prediction of Inter-Branch Angles in RNA Three-Way Junctions from Secondary Structure
Yang H, Qiao N, Zhang B, Shi Y.-Z, and Tan Y.-L.
Biology (2026)
Spatial confinement reshapes the folding of an ion-stabilized DNA with a three-way junction
Wang X., and Shi Y.-Z*.
Chinese Journal of Physics (2026)
gCoSRNA: Generalizable Coaxial Stacking Prediction for RNA Junctions Using Secondary Structure
Li S, Xu Q, Tan, Y.-L, Jiang J, Zhang B, and Shi, Y.-Z.
Biomolecules (2026)
3D structure and stability prediction of DNA with multi-way junctions in ionic solutions
Wang X, and Shi, Y.-Z*.
PLoS Computational Biology (2025)
Branch-specific gene discovery in cell differentiation using multi-omics graph attention
Yin Y., Zhuang L., Wang Y., Shi Y.-Z., and Zhang B*.
PLoS Computational Biology (2025)
KSRV: a Kernel PCA-Based framework for inferring spatial RNA velocity at single-cell resolution
He Y., Jiang J., Qoi H., Shi Y.-Z., and Zhang B.
Frontiers in Genetics (2025)
Advances in RNA contact prediction: a benchmark evaluation of computational methods
Tan Y.-L., Guo C., Xu J.-J, Shi Y.-Z* and Tan Z.-J*.
Communications in Theoretical Physics (2025).
scGRN-Entropy: Inferring cell differentiation trajectories using single-cell data and gene regulation network-based transfer entropy
Sun, R., Cao, W., Li, S.-X., Jiang, J., Shi, Y.-Z., & Zhang, B.-G.
PLoS Computational Biology (2024).
Coarse-grained modeling on sequence-dependent 3D structure size of single-stranded DNA in ion solutions
Yuan, J., Li, S., Tan, Y.-L., Zhang, B.-G. & Shi, Y.-Z*.
CCC 6824–6829 (2024).
ABC2A: A Straightforward and Fast Method for the Accurate Backmapping of RNA Coarse-Grained Models to All-Atom Structures.
Shi, Y.-Z. et al.
Molecules 29, 1244 (2024).
WF-Velo: A New Method for RNA Velocity Analysis Based on Single-Cell Data
Cheng Li, Wenjie Cao, Zihao Zhang, Wenbo Tang, Yazhou Shi, Ben-gong Zhang
DDCLS (2024)
MLBKFD: Probabilistic Model Methods to Infer Pseudo Trajectories from Single-cell Data.
Han Changfeng et al.
Journal of Computational Biophysics and Chemistry (2024).
Computational Modeling of DNA 3D Structures: From Dynamics and Mechanics to Folding.
Mu, Z.-C., Tan, Y.-L., Liu, J., Zhang, B.-G.* & Shi, Y.-Z.*
Molecules 28, 4833 (2023).
cgRNASP: coarse-grained statistical potentials with residue separation for RNA structure evaluation.
Tan, Y.-L., Wang, X., Yu, S., Zhang, B. & Tan, Z.-J.
NAR Genomics and Bioinformatics 5, (2023).
Ab initio predictions for 3D structure and stability of single- and double-stranded DNAs in ion solutions
Zi-Chun Mu, Ya-Lan Tan, Ben-Gong Zhang,J ie Liu, Ya-Zhou Shi
PLoS Comput Biol18(10), e1010501 (2022).
rsRNASP: A residue-separation-based statistical potential for RNA 3D structure evaluation
Ya-Lan Tan, Xunxun Wang, Ya-Zhou Shi, Wenbing Zhang, and Zhi-Jie Tan
Biophysical Journal 21, 142–156 (2022)
RNAStat: An Integrated Tool for Statistical Analysis of RNA 3D Structures.
Guo, Z.-H., Yuan, L., Tan, Y.-L., Zhang, B.-G. & Shi, Y.-Z.
Front. Bioinform. 1, (2022).
Imputation Methods for scRNA Sequencing Data.
Wang, M. et al.
Applied Sciences 12, 10684 (2022).
Salt-Dependent RNA Pseudoknot Stability: Effect of Spatial Confinement.
Feng, C., Tan, Y.-L., Cheng, Y.-X., Shi, Y.-Z. & Tan, Z.-J.
Front. Mol. Biosci. 8, (2021).
SS-RNN: A Strengthened Skip Algorithm for Data Classification Based on Recurrent Neural Networks.
Cao, W., Shi, Y.-Z., Qiu, H. & Zhang, B.
Front. Genet. 12, (2021).
An Integrated Tool for RNA 3D Structure Prediction and Analysis.
Yuan, L., Guo, Z.-H., Cao, W.-J., Luo, Y. & Shi, Y.-Z.
CCDC 4293–4297 (2021).
Predicting 3D structure and stability of RNA pseudoknots in monovalent and divalent ion solutions
Ya-Zhou Shi,Lei Jin,Chen-Jie Feng,Ya-Lan Tan,Zhi-Jie Tan
PLoS Comput Biol14(6) e1006222.(2019)
Structure folding of RNA kissing complexes in salt solutions: predicting 3D structure, stability, and folding pathway.
Jin, L., Feng, C.-J., Tan, Y.-L., Shi, Y.-Z. & Tan, Z.-J.
RNA 25, 1532–1548 (2019).
3D structure stability of the HIV-1 TAR RNA in ion solutions: A coarse-grained model study.
Zhang, B.-G., Qiu, H.-H., Jiang, J., Liu, J. & Shi, Y.-Z.
The Journal of Chemical Physics 151, (2019).
Modeling Structure, Stability, and Flexibility of Double-Stranded RNAs in Salt Solutions
Jin, L., Shi, Y.-Z., Feng, C.-J., Tan, Y.-L. & Tan, Z.-J.
Biophysical Journal 115, 1403–1416 (2018).
Traveling Wave Solutions of Two Nonlinear Wave Equations by (G/G)-Expansion Method
Shi, Y., Li, X. & Zhang, B.
Advances in Mathematical Physics 2018, 8583418 (2018).
3D structure stability of RNA hairpin controlled by loop size
Shi, Y. & Zhang, B.
2017 29th Chinese Control And Decision Conference (CCDC)
Predicting 3D Structure, Flexibility, and Stability of RNA Hairpins in Monovalent and Divalent Ion Solutions.
Shi, Y.-Z., Jin, L., Wang, F.-H., Zhu, X.-L. & Tan, Z.-J.
Biophysical Journal 109, 2654–2665 (2015).
A coarse-grained model with implicit salt for RNAs: Predicting 3D structure, stability and salt effect
Shi, Y.-Z., Wang, F.-H., Wu, Y.-Y. & Tan, Z.-J.
The Journal of Chemical Physics 141, (2014).